Overview
Modern biodiversity research increasingly relies on integrative taxonomic approaches to document, identify, and classify life on Earth.
Traditional morphology based identification has provided the foundation of taxonomy for centuries, but it can be limited when dealing with cryptic species, incomplete specimens, early life stages, or highly diverse groups such as insects, fungi, and microorganisms.
Recent advances in digital identification keys, molecular barcoding, and data integration platforms allow researchers to move beyond single character identification and toward standardized, reproducible, and scalable workflows for documenting biodiversity.
IdentifyLife supports this transition by indexing and connecting identification resources from across the web.
Challenges in Traditional Taxonomic Identification
Morphological Ambiguity and Cryptic Diversity
Many species cannot be reliably distinguished using morphology alone due to:
High phenotypic similarity between species
Intraspecific variation across life stages
Convergent evolution
Degraded or partial specimens
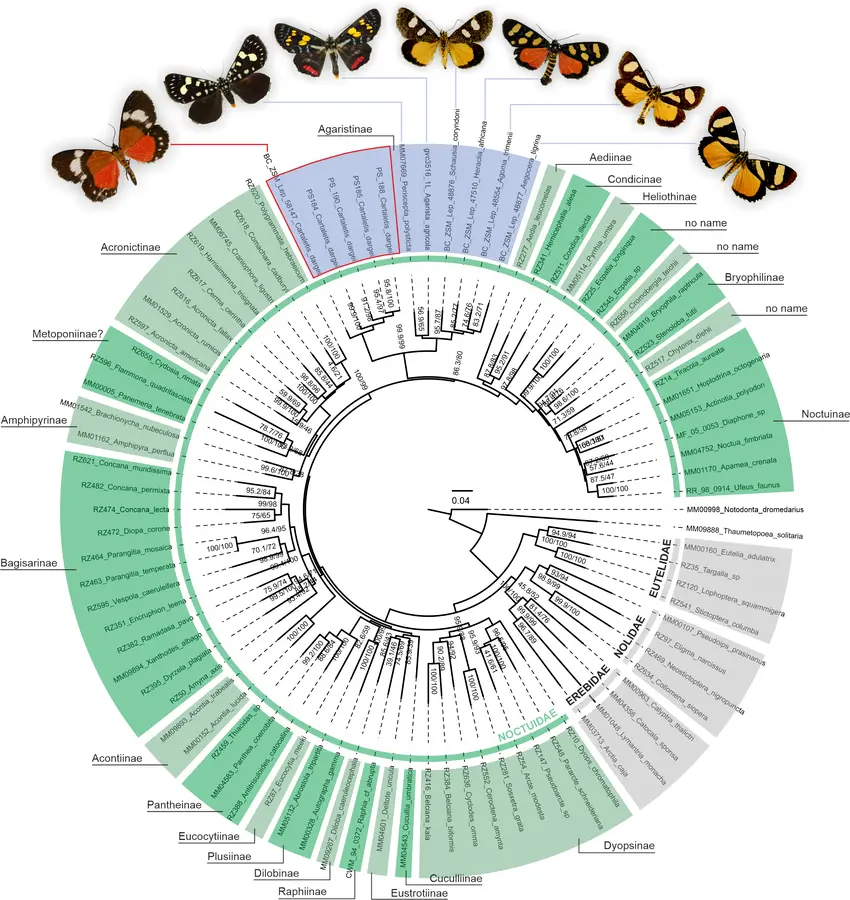
These challenges are particularly pronounced in groups such as arthropods, fungi, algae, and microorganisms, where species richness is high and diagnostic characters may be subtle.
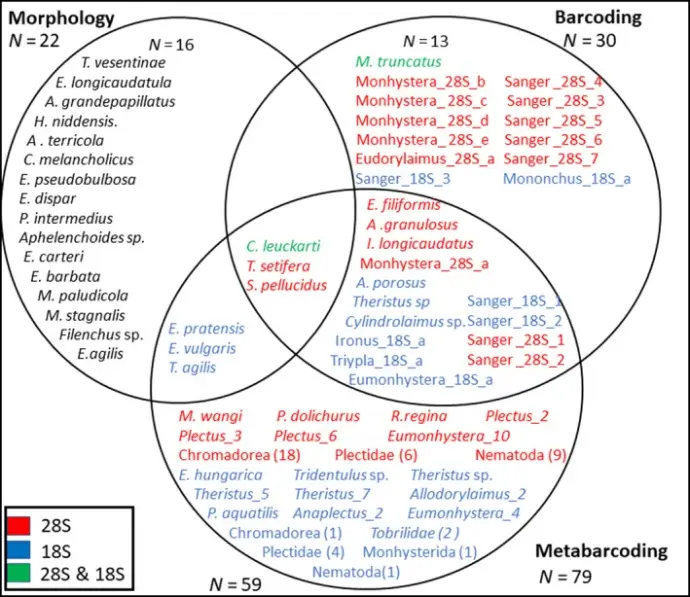
comparison-of-morphological-barcoding-and-metabarcoding.pdf
Identification Keys: Core Tools of Taxonomy
From Printed Keys to Digital Systems
Identification keys remain essential tools in taxonomy. Historically published in books and journals, they now increasingly exist as:
- Dichotomous keys
- Multi-access (polytomous) keys
- Character-based interactive tools
- Image-supported identification guides
Digital keys improve usability, update frequency, and accessibility. IdentifyLife acts as a central index, allowing users to locate appropriate keys across taxa and regions rather than searching fragmented sources.
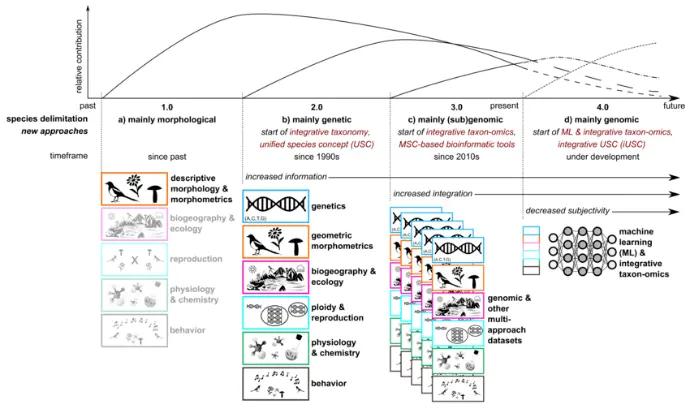
Integrative Taxonomy and Species Delimitation
Combining Multiple Lines of Evidence
Modern taxonomy increasingly adopts an integrative framework that combines:
- Morphological characters
- Molecular markers (DNA barcodes)
- Ecological and geographic data
- Behavioral and chemical traits
This approach strengthens species hypotheses and reduces misidentification, particularly in species complexes.
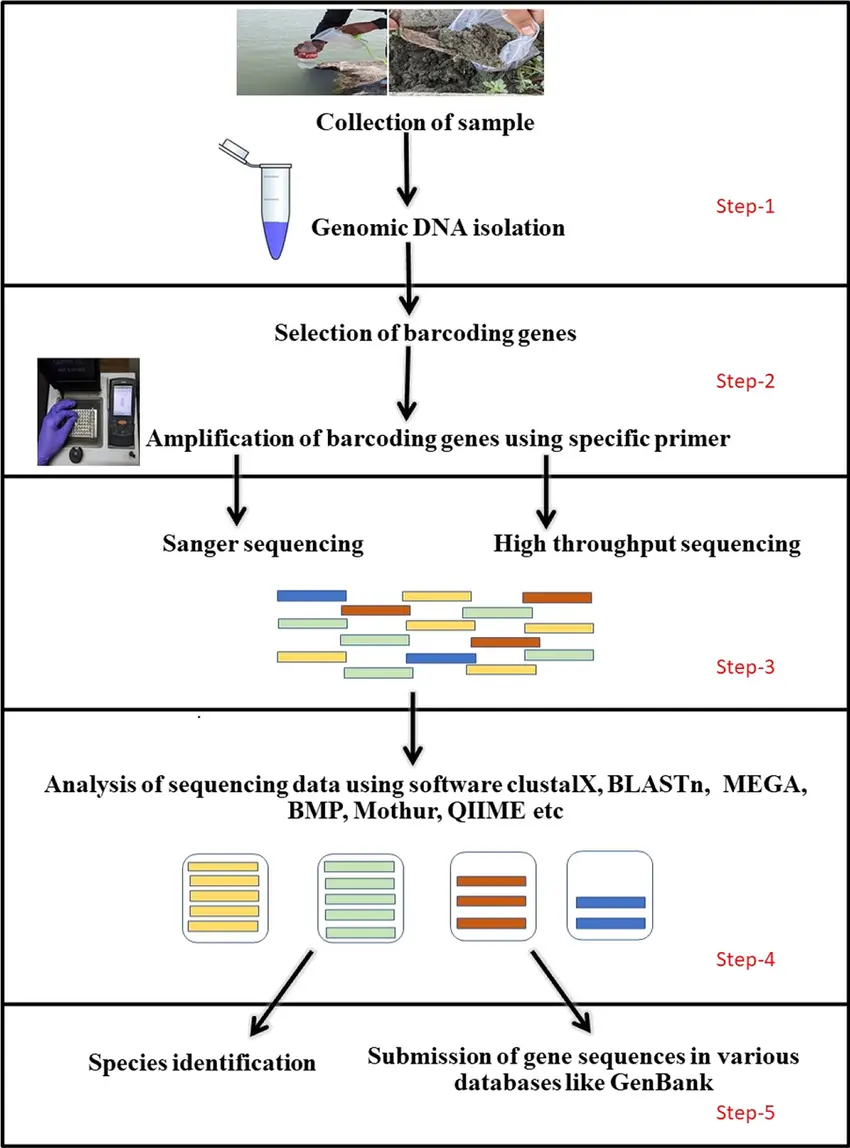
DNA Barcoding and Molecular Identification
Standardized Genetic Markers
DNA barcoding uses short, standardized gene regions to identify species by comparison with reference databases. Common markers include:
- COI for animals
- ITS for fungi
- rbcL and matK for plants
- 16S rRNA for bacteria and archaea
These methods are especially valuable when morphological identification is not possible. IdentifyLife recognizes molecular approaches as complementary tools, aligned with classical taxonomy rather than replacements.
Community and Large-Scale Biodiversity Inventories
All-Taxa Biodiversity Inventories (ATBIs) *A-review-of-convolutional-neural-networks-in-computer-vision.pdf
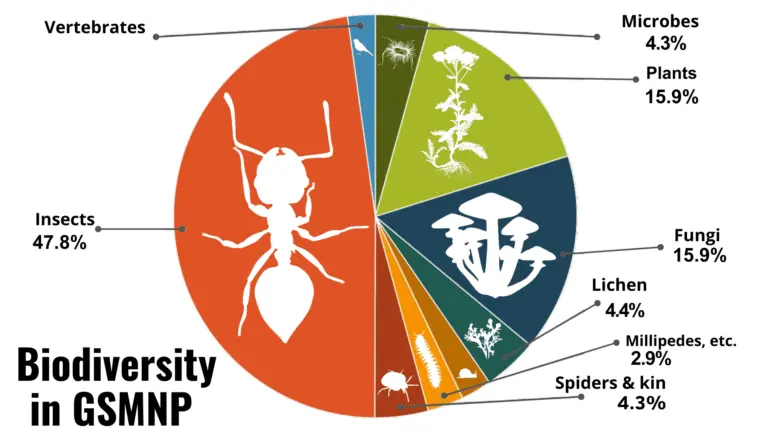
ATBIs aim to document every species within a defined geographic area. One of the most prominent examples is the Great Smoky Mountains ATBI, which integrates taxonomists, molecular methods, and citizen scientists.
Such projects highlight the need for centralized access to identification resources and standardized taxonomic frameworks.
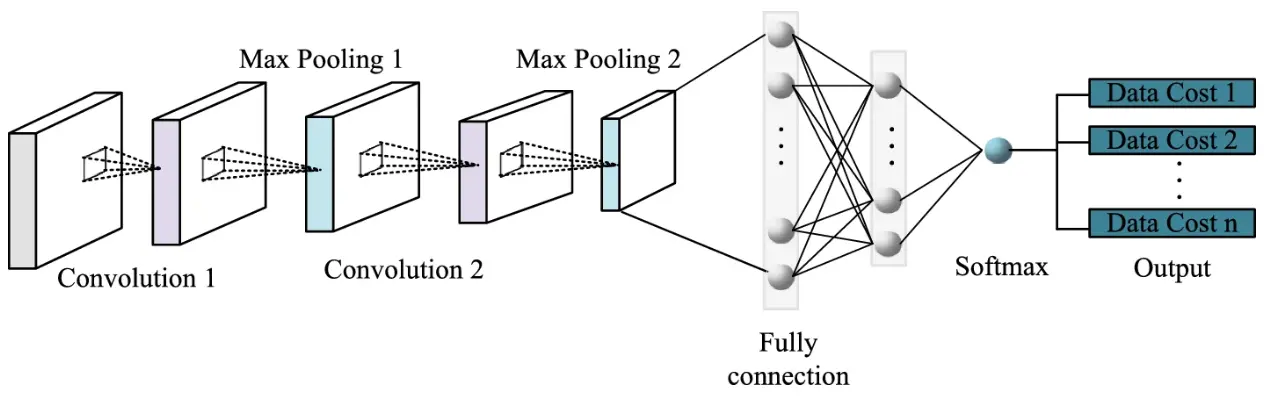
AI-Augmented Identification Approaches
Image-Based Classification
Machine learning, particularly convolutional neural networks (CNNs), has shown strong performance in image-based taxonomic classification when trained on large, curated datasets.
These tools are valuable for:
- Preliminary identification
- Citizen science validation
- Prioritizing specimens for expert review
AI-based systems are most effective when grounded in expert-validated taxonomic frameworks and reliable identification keys.
Linking Identification to Global Taxonomic Frameworks
Standardization and Interoperability
Reliable identification depends on consistent naming and classification. Global taxonomic backbones and species catalogs provide:
- Verified scientific names
- Hierarchical classification
- Synonym resolution
By aligning identification tools with these frameworks, IdentifyLife helps ensure that identification results are interoperable across biodiversity databases, ecological studies, and conservation assessments.
The Role of IdentifyLife in Modern Taxonomy
IdentifyLife serves as a discovery layer rather than a replacement for taxonomic expertise. Its goals include:
- Making identification keys easier to find
- Supporting standardized descriptive data
- Connecting morphology, molecular methods, and taxonomy
- Encouraging reuse of existing high-quality resources
This approach supports researchers, educators, and citizen scientists while respecting the rigor of formal taxonomic science.
Conclusion
Accurate species identification underpins all biodiversity research, from ecosystem monitoring to conservation planning.
As taxonomy evolves toward integrative, data-rich methodologies, platforms that connect identification tools, taxonomic knowledge, and global frameworks become increasingly essential.
By bringing people and identification resources together, IdentifyLife contributes to a more connected, transparent, and collaborative future for documenting life on Earth.